All metrics and proxies distract from science
Scientific research careers worldwide have been profoundly distorted by the focus of nearly all participants upon the metrics and proxies used to evaluate researchers. In particular, informal journal rankings, impact factors, publication counts, H-indices, etc, collectively make or break our careers. Because these metrics are poorly correlated with genuine scientific quality, they have completely deformed the scientific enterprise. The chase for “high-impact” research forces researchers to undertake ridiculously ambitious and unproductive projects whose predictably disappointing results must nevertheless be published, using unjustified hype and over-interpretation. For the same reasons, metrics are also unfair as an evaluation system.
Every supposed metric or proxy of research quality is a distraction. Any time spent comparing metrics is time spent not understanding research. Many people have carefully documented the disconnect between metrics and research quality. Even citation counts, the supposed gold standard, are close to worthless: only a tiny fraction of citations represent serious evaluation or validation, whereas the great majority represent nothing more than lazy copying and fashion-following. Two anecdotes demonstrate just how worthless citation counts can be:
- Some of the most highly-cited researchers in the world have been exposed as gaming the citation system and as charlatans.
- Papers with thousands of citations turn out to be false; obviously not one of those citations represented a serious validation.
There is currently no way to distinguish a genuine citation from a gamed one, and currently nobody identifies which citations represent a genuine replication of previous work. In sum, interpretation of citations is meaningless in the extreme.
Many involved in research have realised the problems of metrics, and there are some efforts to change the system, including the San Francisco Declaration On Research Assessment (DORA) alluded to in the title. However, although large numbers of organisations have now signed the declaration, many people are unconvinced that it will be politically and practically possible to replace metrics with more genuine evaluation processes. It is true that abandoning metrics will force us to change our procedures, but that would surely be a good thing. How might the new system work? Below are some suggestions for non-metric evaluations, drawn from my experience as a researcher, as a member of (CNRS) recruitment committees and as an organiser of PubPeer. (Working at PubPeer soon disabuses one of any notion of a reliable connection between scientific quality and any formal or informal metric of excellence.) The suggestions are mostly written from the viewpoint of a recruitment or promotion committee, but would also be relevant in reviews of grant and fellowship applications or performance reviews. Few if any of my suggestions are original, I’m just adding my brick to the wall. I’ve been particularly influenced by David Colquhoun who has written repeatedly about this.
Evaluation is difficult, so do it infrequently
In a typical recruitment session at the CNRS, I would be asked in a short time to evaluate in detail 10-15 candidates. All of the candidates were neuroscientists and my applications were those most appropriate for my expertise. Although with some effort I felt I could understand what those candidates were doing and find (I believe) reasonably intelligent questions for most of them, what I found extraordinarily difficult was to situate the candidates within their fields of research, because that required intimate knowledge of their specific subfield. In general, I was only comfortable in making an overall quality/impact judgement for a handful (no more than 25%) of my candidates. Obviously, I was mostly at sea for the candidates evaluated by my colleagues in the committee. The system was unavoidably superficial, despite the best efforts of the committee members.
If we (rightly) forbid metrics and instead base evaluation upon an in-depth understanding of a researcher’s work and their position in their field, my experience illustrates how this will require expertise and time, and therefore be expensive. The inescapable consequence is that one should do evaluation as infrequently as possible, and only when it is absolutely necessary. One should also only undertake it when motivated to do it properly, which is when something important is at stake. I would suggest performing such evaluations at no more than a few major career stages – recruitment, significant promotions. In contrast, I would simply dispense with annual evaluations, which are mostly ineffective, time-wasting micro-management. I also think we shouldn’t be hiring evaluators to count grant money; not getting a grant is its own feedback and punishment, so re-evaluating that seems like a double jeopardy.
Is their best good enough?
One of the hardest stages at which to give up metrics is the triage stage. There might be a hundred candidates to screen. Nobody can read and understand the full scientific output and context of that many applicants. Often this is still done by a formula along the lines of “at least one first-author paper in a journal with an impact factor greater than 10″… What I suggest is to focus instead on a brief (one page?) description of the candidate’s most important research contribution, selected by them and described in their own words. The logic here is simply that if their best is not good enough, then it’s not worth continuing with them. We see in this approach the importance of making detailed author contribution statements in papers; without these it may be impossible to verify objectively a candidate’s contribution in today’s highly collaborative research.
Focusing on a candidate’s best work naturally favours quality over quantity. This criterion of first judging their best work should be carried right through the evaluation process. To repeat: why select somebody whose best is not good enough?
Is their worst good enough?
I doubt I’m alone in feeling swamped by low-quality publications. Indeed, experience with PubPeer has shown that a significant fraction of papers are worse than low quality – the fruit of complete incompetence or frank misconduct. I believe that until publishing something wilfully erroneous or fraudulent becomes a career negative, researchers will not concentrate sufficiently on producing high-quality work. Unfortunately, publication even of outright rubbish is still generally a career positive, especially if it is in a glamour journal (there are many examples of this, a few are detailed elsewhere on this blog and many more are discussed on PubPeer; you probably also know your own examples).
I therefore propose to downgrade/eliminate candidates who have published work of low quality or worse. Naturally, it will rarely be productive to ask a candidate to identify their worst work! However, searching the PubPeer database, journal commenting systems, other online discussions and possibly talking with colleagues may on occasion throw up a lead, in which case it should be followed and taken seriously. If there are no leads, a couple of papers could be chosen for inspection at random. Again, we see the importance of detailed author contribution statements, to protect innocent co-authors in collaborative papers. However, with negative contributions as for positive ones, sometimes we must take a decision on the basis of incomplete information (“did the fourth co-first author really do all of the work?”).
Are they competent?
Something that is nearly absent from modern recruitment is the evaluation of technical competence; it appears that the only skill that matters is being able to elbow one’s way into (a good position on) an author list. It is very worthwhile to ask the candidate explicitly to list and document the skills and techniques they have acquired. That list and the Methods section of a recent article could then form the basis of a revealing discussion. It sometimes becomes apparent that a candidate has not mastered or is even unfamiliar with the techniques they reportedly applied. The appropriate list of useful skills is obviously context dependent. Personally, I feel that important but undervalued general skills include quantitative ability, programming and statistical understanding.
On a practical level, it is often difficult to verify the ability and contributions of one author amongst many, with the only opportunity occurring during a sometimes short audition (CNRS auditions are ridiculously brief). In previous times, young researchers were encouraged to publish at least one paper alone, to prove unequivocally what they could do. This should of course be noted if the case arises, but it is much less common today.
With the aim of forming a more holistic opinion of a candidate’s scientific reasoning, rich insight is sometimes available if they write a blog or contribute to post-publication commentary, because these are often very personal, individual contributions. Finally, additional information can sometimes be obtained from social media accounts (including mouth-watering recipes from Provence). It is therefore worth checking for such activities.
Does their work replicate?
Instead of counting papers and citations, why not check whether anybody has replicated the candidate’s work? Indeed, ask them to list/describe those replications. A replication is a heavyweight validation. If somebody both found the work of sufficient interest to replicate and was indeed able to obtain the same results, that speaks both to the interest of the work in the field and to its correctness. Obviously, independent replications are greatly to be preferred. Unfortunately, for early-career researchers, there may not have been enough time for any replications to have been performed, but this could be a very useful criterion for more advanced researchers.
Conversely, unresolved failures of replication should probably be considered as a strong negative indicator and merit detailed investigation. An impression I’ve gained from involvement in PubPeer is that unresolved failures to replicate are strong predictors of problematic research. Although it’s just an anecdote, I found Jeffrey Flier’s retrospective on involvement in three misconduct cases to be revealing. At the time he hired Pierro Anversa and handed him the keys to Harvard, a colossal mistake, the only visible red flag seems to have been that at least two researchers publicly disputed his work.
In addition to the results, the manner in which an author reacts to a replication attempt (and then to its eventual failure) could also be a criterion. Were they helpful or obstructive? Did they share methods, materials, data, code?
Are they good mentors?
Although some institutions may willingly hire a slave-driver and turn a blind eye to exploitation of early-career researchers as long as grant money flows and papers are published in glamour journals, let’s assume that most selection committees would prefer to reward conscientious mentors who care about the current and future well-being of their lab members. A radical approach to ensuring this would be to request letters of recommendation about the candidate from those they have previously mentored. Apparently this is part of the tenure process at Stanford, although there was scepticism as to whether such information had ever led to tenure being refused. In any case, such an approach could generate some systemic improvements. It seems plausible that PIs would treat their lab members a lot more fairly if they knew that they would in the future be writing recommendation letters. There is also a clear if cynical argument of enlightened self-interest for institutions to request such letters: they may point to a pattern of bullying, harassment or misconduct. Indeed, researchers with such histories tend to move frequently to keep ahead of trouble, with each previous institution keeping silent to rid themselves of the problem. It’s always better to know about such problems before hiring somebody.
Such recommendation letters may indicate the conditions under which young researchers leave a lab. One thing to watch out for is the PI who considers previous lab members to be potential competitors or who makes intrusive efforts to regulate their choice of research subject.
Favour open science and reproducibility
If you want the research environment to evolve, why not select for virtuous behaviour explicitly? Even better, include these criteria in your announcement, to send a signal. Here is a brief list of some items that might be positively evaluated:
- public data sharing (candidates could list papers that reused their data)
- preregistration of studies
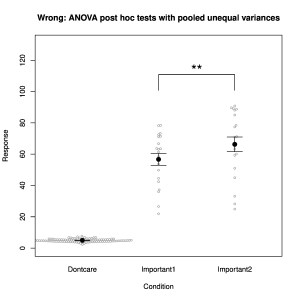
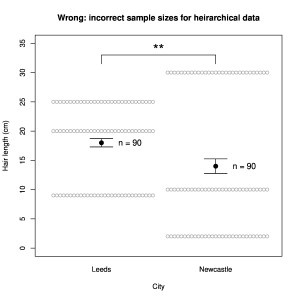
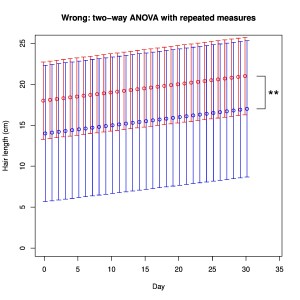
- use of proper sample sizes
- release of open source code
- preprints
- contribution to post-publication peer review
Candidates can’t change the system?
If you are on a recruitment or grant committee, you can of course help design the application procedure according to ideas such as those listed above. But if you are a candidate, it is rarely open to you to dictate the application procedure. However, the situation may not be as hopeless as you imagine. With a bit of care, it will often be possible to insert information corresponding to the items above into your application. Even if it has not been explicitly requested, evaluators are likely to notice this information and will then naturally compare your strong points with those of other candidates. They may also remember to redesign their application form the next time around.
In the end, nearly all evaluations, reviews and selections are carried out by researchers. We always have some room to stretch the criteria in the direction we feel rewards high-quality research. The system probably won’t change if nobody tries to change it, but it might if we try together.