A high-profile article in Nature Materials reports an ultra-ultra-sensitive, enzyme-linked assay. According to the paper, 0.0375 molecules of enzyme produce the maximum assay signal. No plausible mechanism has been offered for this sub-Avogadro performance. The authors make the kinetically absurd argument that the sensitivity of the assay is increased, even at concentrations near the single-molecule limit, by the presence of a competing reaction that reduces signal. This topsy-turvy, less-is-more mechanism is dubbed “inverse sensitivity”.
The assay is a bit complicated [1]. In brief, in the presence of the analyte antigen, the enzyme glucose-oxidase (GOx) is retained and then produces H2O2, which subsequently reduces silver ions that are deposited on gold nanostars (nanoparticles) and detected via a blueshift of the solution absorbance peak. The competing reaction supposed to increase sensitivity is nucleation of silver particles elsewhere than on the gold nanostars.
Public discussion of the paper was initiated on PubPeer by a commenter pointing out: i) a huge discrepancy between the amounts of silver likely to be produced and that necessary to generate the observed blueshift, and ii) that one of the reactions appeared to detect less than a single molecule of enzyme.
I joined the discussion (as Peer 2), confirming these problems. My calculations suggested that the likely silver production and that required to account for the observations were inconsistent by a factor of at least 1011. Additionally, the reported sensitivity of the assay required an exceptionally stringent absence of silver ion reduction in the absence of H2O2, otherwise the small signals caused by the analyte would be completely swamped. However, the reaction mixture is essentially the well known “silver mirror reaction”, in which glucose reduces bulk quantities of silver.
The authors did not enter the discussion on PubPeer or supply any scientific response when contacted directly. The concerns about the article were written up and communicated to the journal editors. The analysis was also eventually posted as a preprint.
After a confidential and obscure process lasting more than four years, the journal published in short succession an uninteresting Corrigendum and a much more interesting Addendum, the main subject of this post. The Addendum is supposed to add an explanation that was missing from the original article without otherwise changing its conclusions. It does much more than that:
- it offers a (rather implausible) new reaction mechanism that demolishes the central role for the titular “inverse sensitivity”
- it throws one whole figure under the bus
- it creates new inconsistencies
First the new mechanism. The authors suggest that most of the silver deposited on the nanostars does come from reduction by glucose or other non-analyte-related routes. (Interestingly, the Addendum lists a large number of possible reducing mechanisms, most of which were not mentioned in the public discussion, raising the possibility that other complaints were received by the journal or the authors.) However, in order to retain assay selectivity, the authors suggest that this bulk deposition cannot occur spontaneously, but can only amplify “seed” deposition induced by the “stronger reducing agent” H2O2. This seems like a strange assertion on its face, since descriptions of the silver mirror reaction present the reduction as a reaction in solution without mention of a need for any special physical substrate. However, I’m not a chemist, so I’ll leave the evaluation of the plausibility of this step of the reaction to experts. (The Addendum offers complex but unverified mechanisms for selective deposition; see the Appendix to this post.)
More to the point, the authors now explain the extreme sensitivity of their assay through this amplification mechanism: “… a small concentration of enzyme-generated hydrogen peroxide can trigger the formation of a silver coating around gold nanostars. This phenomenon is likely responsible for the ultralow limit of detection reported in the Letter.” So there is no role for inverse sensitivity in explaining the reaction sensitivity at low analyte concentrations. For sure, the authors still talk about inverse sensitivity in the Addendum, but it only reduces the sensitivity of the assay, at higher analyte concentrations. The elimination of inverse sensitivity as the (nonsensical) source of the high sensitivity is a major change. As the following quotes recall, “inverse sensitivity” is the central point of the paper (emphasis added in each citation):
Title: “Plasmonic nanosensors with inverse sensitivity by means of enzyme-guided crystal growth”
Abstract: “Lowering the limit of detection is key to the design of sensors needed for food safety regulations, environmental policies and the diagnosis of severe diseases. However, because conventional transducers generate a signal that is directly proportional to the concentration of the target molecule [this is the law of mass action], ultralow concentrations of the molecule result in variations in the physical properties of the sensor that are tiny, and therefore difficult to detect with confidence. Here we present a signal-generation mechanism that redefines the limit of detection of nanoparticle sensors by inducing a signal that is larger when the target molecule is less concentrated [they really did write that]. The key step to achieve this inverse sensitivity is to use an enzyme that controls the rate of nucleation of silver nanocrystals on plasmonic transducers. We demonstrate the outstanding sensitivity and robustness of this approach by detecting the cancer biomarker prostate-specific antigen down to 10(-18) g ml(-1) (4 × 10(-20) M) in whole serum.”
Cover page caption: “Conventional sensors generate a signal that is directly proportional to the concentration of the target molecule [the law of mass action]. Now, by means of an enzyme that controls the growth of silver nanocrystals on plasmonic transducers, a nanosensor with sensitivity that is inversely proportional to concentration, and can detect ultralow concentrations of the cancer biomarker prostate-specific antigen in whole serum, is demonstrated.”
According to COPE guidelines (pdf): “Journal editors should consider retracting a publication if they have clear evidence that the findings are unreliable, either as a result of misconduct (e.g. data fabrication) or honest error (e.g. miscalculation or experimental error).” The original version of the paper found that inverse sensitivity was the mechanism for the ultra-low detection threshold, a finding that would seem rather unreliable now that it has been discounted in favour of the amplification mechanism by the authors themselves! (Disclaimer: maybe you need to be a professional editor to interpret COPE retraction guidelines correctly.)
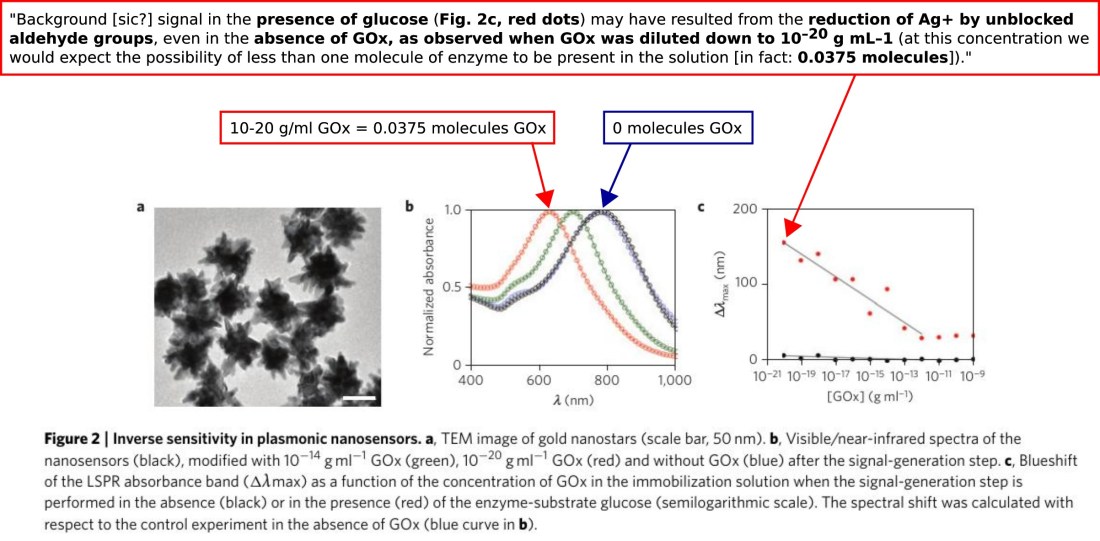 The interpretation of Fig. 2 has changed significantly. In panel b, we see that huge spectral shifts occur when the only change is from zero GOx to much less than one molecule of GOx. A mechanism for this homoeopathic action has still not been supplied. In panel c, the maximum signal is now attributed to a contamination. © Springer Nature, reproduced with permission.
The interpretation of Fig. 2 has changed significantly. In panel b, we see that huge spectral shifts occur when the only change is from zero GOx to much less than one molecule of GOx. A mechanism for this homoeopathic action has still not been supplied. In panel c, the maximum signal is now attributed to a contamination. © Springer Nature, reproduced with permission.Further bombshells in the Addendum concern the data of Fig. 2 (above). The authors confirm that the reaction volume was 1 ml, which in turn confirms that maximum signal was observed when the assay shown in Fig. 2c contained 0.0375 molecules of glucose oxidase. In other words, none. The authors bow to the inevitable and now acknowledge this, albeit in a mealy-mouthed way: “… when GOx was diluted down to 10−20 g mL−1 (at this concentration we would expect the possibility of less than one molecule of enzyme to be present in the solution)”. Furthermore, if the maximum signal in Fig. 2c is not the result of GOx, the authors cannot escape offering another explanation. It is suggested to arise from a contamination: “… signal in the presence of glucose (Fig. 2c, red dots) may have resulted from the reduction of Ag+ by unblocked aldehyde groups, even in the absence of GOx, as observed when GOx was diluted down to 10−20 g mL−1.” The authors don’t present any new experiments to establish their tentative reinterpretation of Fig. 2, so it is now worthless. Thus, they have implicitly conceded that the maximum signal in a key figure is entirely due to a contamination. Why that should be considered an “additional explanation” worthy of an Addendum rather than a Corrigendum (at best) is unclear.
That still leaves the excruciatingly awkward question of what really happened in Fig. 2b, which is still not addressed in the Addendum. The authors acknowledge (see above) that at 10−20 g mL−1 there is no GOx and suggest that signal arises from a contaminating reductant. If there is no GOx, removing it should make no difference. Yet, in Fig. 2b, removal of this ultra-ultra-low (zero) concentration of GOx is reported to generate the maximum spectral shift! How did the authors, editors and referees convince themselves that an assay yielding maximum signal in response to 0.04 molecules of GOx was so banal that it was unnecessary to supply a mechanism? Developing an explanation was left as an exercise for the reader?
In summary, the journal editors took more than four years to produce this fabulously indulgent Addendum. It offers a highly complex, unsubstantiated mechanism in replacement of the physically implausible (impossible) one previously proposed. An entire figure is shown to be unreliable and some of the results—notably the massive effects of 0.04 molecules of GOx—still cannot be explained by any mechanism so far proposed (or known to mankind*). The foot-dragging in handling this case was one of the drivers behind a blog post criticising the Nature editorial stance on refutations, although the publication of the Addendum should be acknowledged. The Nature Materials editorial team are collectively responsible for this case, but the fact that Pep Pàmies (now at Nature Biomedical Engineering) was copied into some correspondence raises the possibility that he had a specific involvement. Mikael Käll wrote a News and Views piece about the original article, which suggests he may have refereed the paper.
I would welcome discussion, here or on the PubPeer thread.
* Unless one counts a mix-up in the concentration calculations.
Update: Matter arising
Nature Publishing Group have launched a new publication format across many of their journals: Matters Arising. This seems quite a promising channel for post-publication discussion. As the editorial says, “This should allow debate on published papers in the journals’ online pages”. I wondered whether, with this new format, the editors of this paper would be more welcoming of “debate” than previously. I submitted a focused Matter Arising (pdf). I pointed out that the inverse sensitivity assay can respond to identical solutions with either maximum or minimum signal. I mildly suggested that without an explanation such variance might, potentially, undermine the confidence of readers in the assay. In my cover letter I expressed my desire for this single point to be clarified.
I needn’t have got my hopes up. The submission was desk rejected without review. I’m not allowed to quote the editor’s confidential, copyrighted decision, but the essence of it is that there must be reasonable explanation somewhere. What or where that explanation is remains their special secret.
Reference
[1] Rodríguez-Lorenzo L, de la Rica R, Álvarez-Puebla RA, Liz-Marzán LM, Stevens MM. (2012) Plasmonic nanosensors with inverse sensitivity by means of enzyme-guided crystal growth. Nat Mater. 11:604-7. doi: 10.1038/nmat3337.
Appendix
Mechanism of silver deposition
The authors’ new amplification mechanism requires that no silver at all be deposited on the nanostars in the absence of H2O2 (produced by GOx). This in turn requires that the relatively minor modification of the conditions of the classic silver mirror reaction—originally optimised to deposit silver in bulk quantities—prevents any deposition of silver at all. The experimental change is to add 5 mM MES at pH6 to a solution containing 40 mM NH3. I would expect only a modest change of pH—could it really extinguish all silver deposition? A key control is therefore to show no signal is observed without the GOx. This is inferred from Fig. 4. However, for labs with access to gold nanoparticles and the absorbance measurement, it might be relatively simple to verify these mechanisms by mixing MES, NH3, AgNO3 and H2O2. Anybody potentially interested in carrying out such experiments is encouraged to contact me. (NB. I tried to prepare the reaction mix without nanoparticles and obtained a pH of 10.5, not the reported 9, so it would be worth checking this parameter.)
Stochastic noise
Amongst the original criticisms (including from me) was the absence in the reported signal of stochastic noise that might be expected from a process detecting a small number of molecules. In the Addendum, the authors point out that the variation of the signal depends on the shape of the dose-response curve. This is correct and my analysis on this point was wrong (or at least incomplete). However, an interesting question is raised. The authors go to great length to attach GOx molecules to the nanostars and lead (or at least allow) readers to believe that the H2O2 produced by an attached GOx molecule will act on the local nanostar. This is in fact unlikely to be the case, for several reasons outlined in the preprint. Moreover, if it were the case, then only a tiny number of nanostars would be coated with silver and the resulting tiny signal would demonstrate Poisson statistics. In summary, there is no kinetic advantage to attaching anything to the nanostars.